This project develops interpretable model and network selection methods for high-dimensional data. The motivating problem is that many predictive models can achieve similar accuracy, while telling very different scientific stories. Instead of selecting a single apparently optimal model, this line of work studies sets of models with comparable predictive performance and uses their structure to reveal stable, ambiguous, or context-dependent relationships among variables.
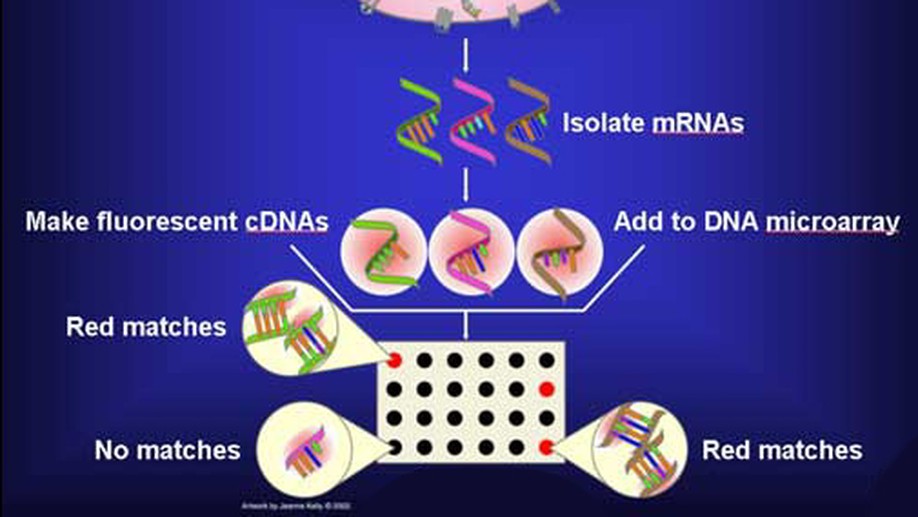
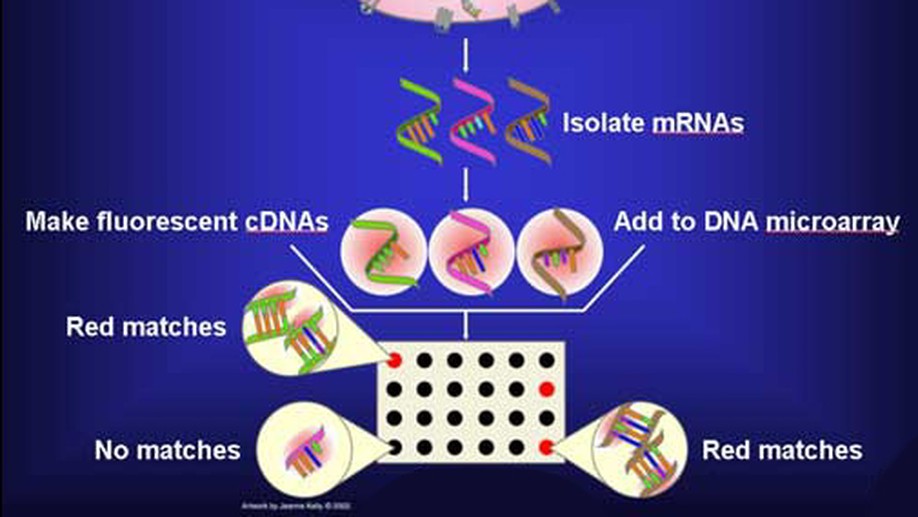
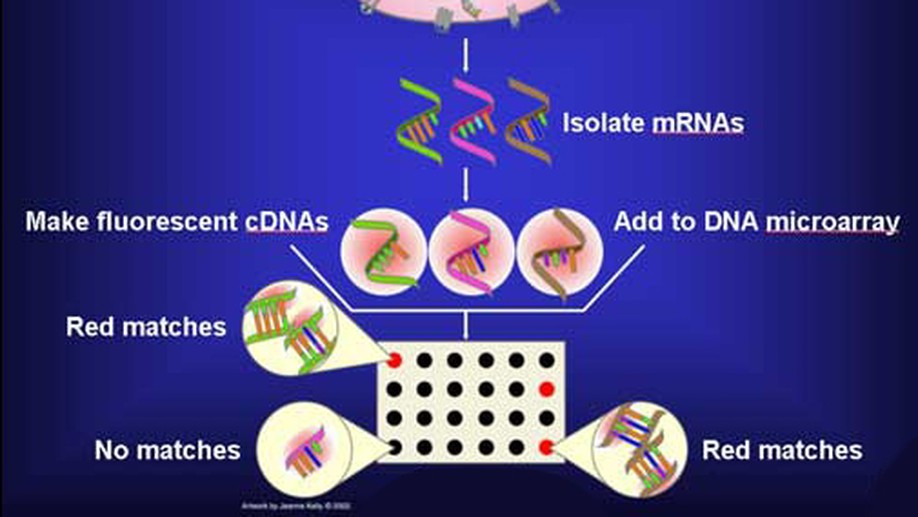
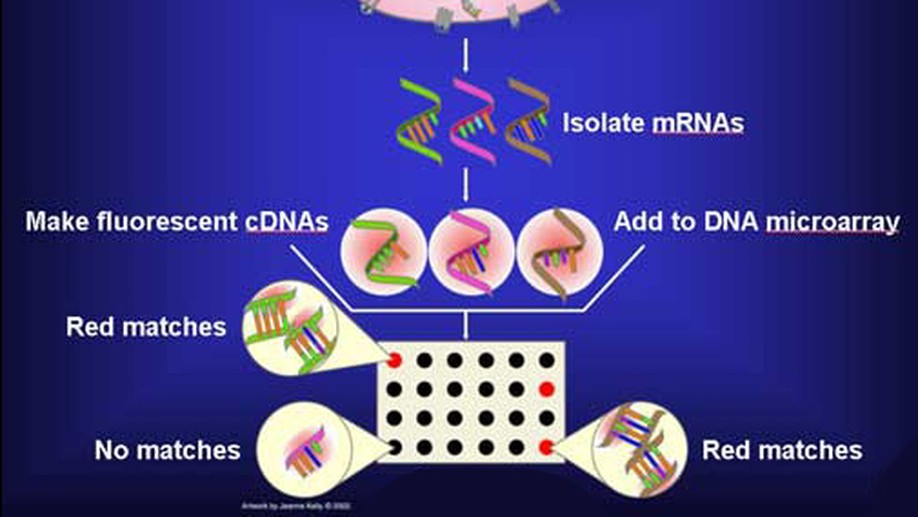
The SWAG framework is central to this direction. It is a wrapper method for sparse learning that emphasizes both predictive ability and interpretability. In applications to genomics and biomedical data, this perspective is especially useful because correlated biomarkers can have competing or even antagonistic roles. The resulting selected model sets can be interpreted as networks, making it possible to identify variables that are consistently active, variables whose effects depend on their context, and variables whose apparent importance is unstable across equivalent models.
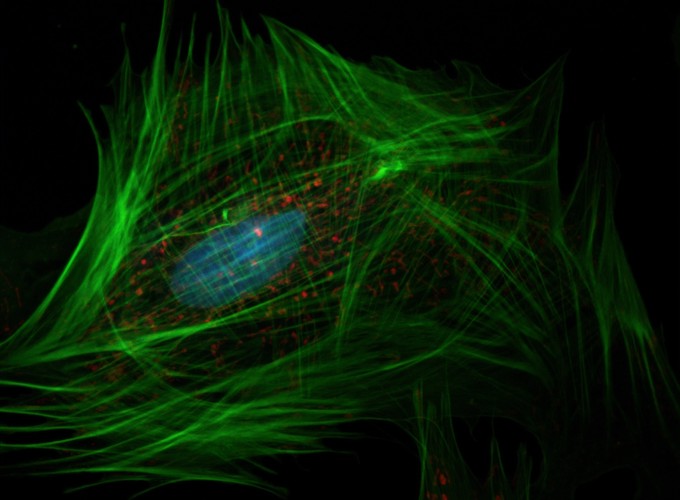
The project has been applied to gene selection, inflammatory bowel disease, melanoma, breast cancer microRNAs, neuroimaging, stellar blend classification, sleep epigenetics, and medical decision support. Across these applications, the goal is the same: build statistical learning tools that are not only accurate, but also scientifically interpretable.
Selected related publications include:
- Swag: A Wrapper Method for Sparse Learning
- A Predictive Based Regression Algorithm for Gene Network Selection
- A Paradigmatic Regression Algorithm for Gene Selection Problems
- Evidence of antagonistic predictive effects of miRNAs in breast cancer cohorts through data-driven networks
- Is Nonmetastatic Cutaneous Melanoma Predictable Through Genomic Biomarkers?
- Stellar Blend Image Classification Using Computationally Efficient Gaussian Processes
- A Multi-Model Framework to Explore ADHD Diagnosis From Neuroimaging Data
- Epigenetic Impact of Sleep Timing in Children: Novel DNA Methylation Signatures via SWAG Analysis